Jupyter Notebook (or Lab) is great for prototyping but not really suited for writing good code. I love Notebooks for trying out new things, plotting, documenting my research, and as an educational tool. However, they don't help you like an IDE with, for instance, code linting and refactoring. Notebooks written by data scientist are notorious for being unreadable, unreproducible and full of bugs; how can we get them to write better code?
One solution for writing less terrible code in Notebooks, is to only use an IDE and write no code in Notebooks. But wouldn't it be great if you could have both the assistance of an IDE and the interactivity of a Notebook?
TL;DR
- Move your code from your Notebook into a Python package.
- Install the package in your virtual environment in development mode.
- Use the
%autoreloadmagic to automatically re-import updated code.
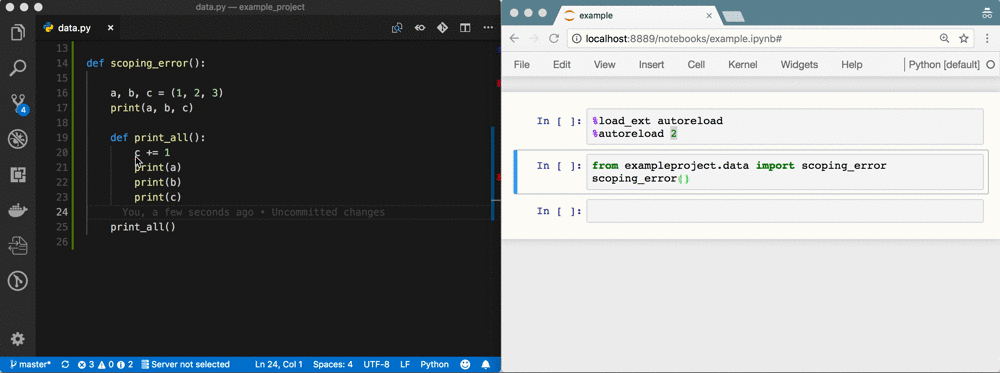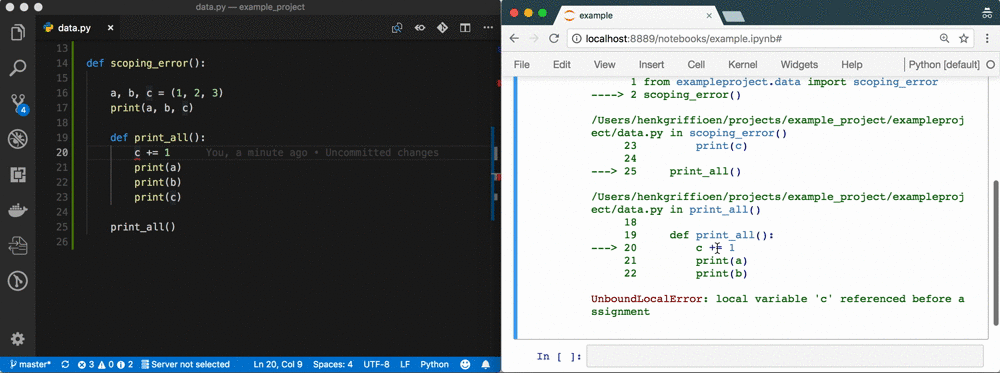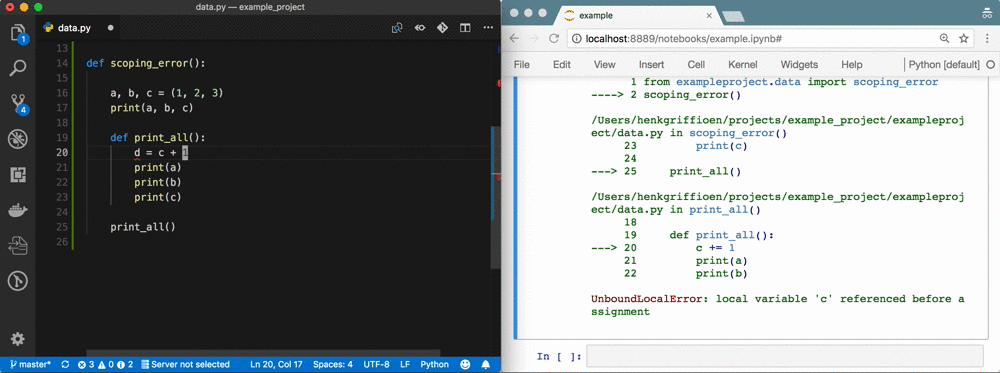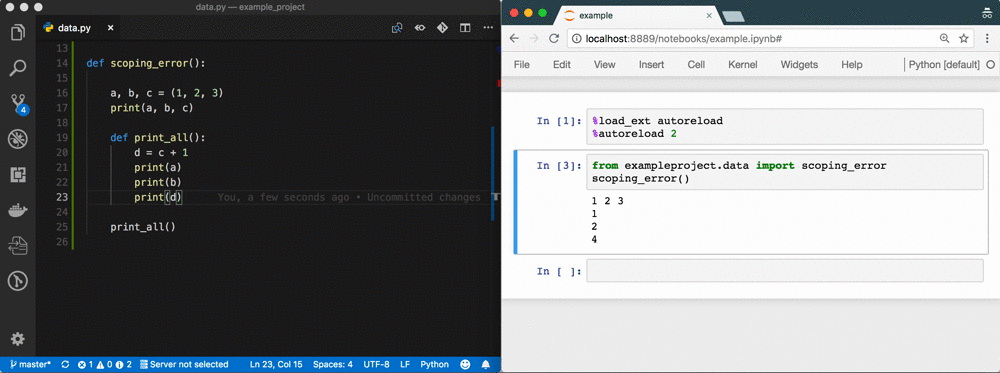
The problem
A week of data sciencing often results in pages of spaghetti code in a Notebook called Untitled4.ipynb. Code has been copy-pasted throughout the Notebook; abbreviations are used for variable names; variables are used that don't exist anymore or are from the wrong scope; there's no reproducibility because of a non-linear flow 1; execution count is in the thousands; etc. It's a mess.
Dream data science/engineering conference agenda pic.twitter.com/qnIY3qicRT
— Vicki Boykis (@vboykis) 16 november 2017
We can fix number 9!
Why does this happen? Data science is often an exploratory process to validate an idea and writing code is one of the science tools. Compare this to engineering, where writing high quality code is one of the key requirements - the exact implementation may also be unknown, but the code has to be solid.
In a perfect world, data scientists would build models and of course create high quality code, but the process and the tools used don't always help.
1. From Notebook to package
To move beyond prototyping, your collection of Notebooks should be transformed into a neat, well-structured Python package. My projects generally look something like this:
example_project/
├── notebooks/ <-- Jupyter notebooks.
└── Untitled4.ipynb
├── exampleproject/ <-- Python package with source code.
└── __init__.py <-- Make the folder a package.
└── example.py <-- Example module in the package.
└── setup.py <-- Install and distribute your module.Prototyping initially happens in the notebooks/ folders, but most logic moves to the package exampleproject as ideas get validated and the overall structure is more clear.
An IDE like PyCharm or Visual Studio Code can assist with developing code. These editors warn upfront when you make a mistake and help you with code style, refactoring, etc. Notebooks, however, are not that intelligent and only warn you if do something really weird with syntax, or complain after running code.
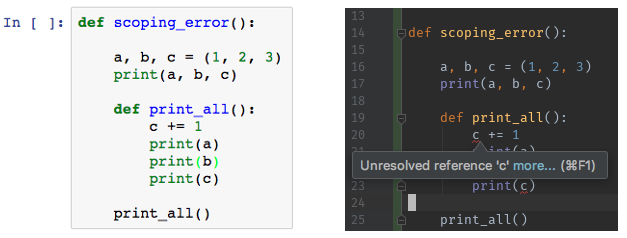
IDE's really shine when your code is structured as a Python package. That is the structure they expect and understand, so you will really get the benefits if your organize it properly2.
2. Using your package in a Notebook
Editing code in an IDE is nice, but you also want to call your functions in a Notebook and test if everything works. To make your package available to your Notebook, activate the conda virtual environment example_venv and install the package in folder exampleproject/:
$ pwd
~/example_project
$ conda activate example_venv
$ pip install --editable .
running develop
...
Finished processing dependencies for exampleproject==0.0.1The setup.py contains instructions for pip to install the package in example_venv3. Making it available by installing saves you from doing scary things like sys.path.append('../'). Option --editable allows you to edit your code without re-installing the package to get the changes.
You can now import your package in your Notebook:
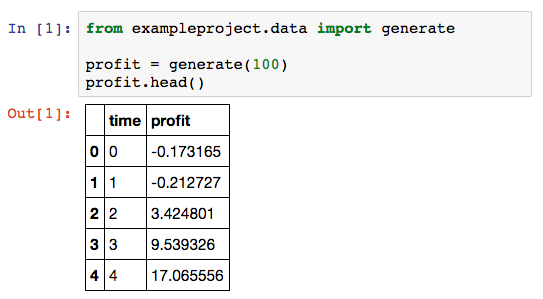
One disadvantage of using a package is that you need to restart the kernel if you change the library code. Although you have now gained the power of an IDE, you are missing out on the interactivity of the Notebook. We need just one more step to fix that.
3. The %autoreload magic
How can you get the interactivity back and get our changes immediately in our Notebook? Add %autoreload at the top of your Notebook:
%load_ext autoreload # Load the extension %autoreload 2 # Autoreload all modules
%autoreload is a Jupyter extension that reloads modules before executing your code. Functions and classes loaded in notebooks get their functionality updated every time you execute a cell. This means that when new code is saved in the editor, the changes are immediately loaded in your Notebook if you run a cell4.
Using %autoreload bridges the gap between Notebook and IDE 5. You gain all the benefits of an IDE, but you're still as flexible as before! See the GIF at the top as an example.
Conclusion
Notebooks don't have the functionality of IDE's, but that does not mean we can not use them while developing code! The package-and-autoreload-flow can help Data Scientists to transform prototype code into production code more easily. Check out my repo for an example of a structure for a data science project, and my previous blog for basic components of project.
Props to Diederik for reviewing this blog!
Learn Spark or Python in just one day
Develop Your Data Science Capabilities. **Online**, instructor-led on 23 or 26 March 2020, 09:00 - 17:00 CET.
Improve your Python skills, learn from the experts!
At GoDataDriven we offer a host of Python courses from beginner to expert, taught by the very best professionals in the field. Join us and level up your Python game:
- Python Essentials - Great if you are just starting with Python.
- Data Science with Python Foundation - Want to make the step up from data analysis and visualization to true data science? This is the right course.
- Advanced Data Science with Python - Learn to productionize your models like a pro and use Python for machine learning.
- See this Twitter thread by Jake Vanderplas and this magic to generate a linear history. ↩ - For instance, the project root there's only one Python package in the project folder and this folder does not contain an
__init__.py. ↩
- Or use python setup.py develop --no-deps (check the docs). ↩
- If you don't feel like adding %autoreload to every Notebook, add it to your IPython profile. ↩
- Don't use %autoreload in your production code though! All kind of nasty things may happen. ↩Written by
Henk Griffioen
Our Ideas
Explore More Blogs
Contact




